Integrated Analysis of Cytoskeleton-Associated lncRNAs and Their Regulatory Networks in Mouse Oocyte Maturation
DOI:
https://doi.org/10.22452/mjs.vol44sp1.11Keywords:
Cytoskeletal proteins, Long non-coding RNAs, Mouse oocyte development, ceRNA, reproductive healthAbstract
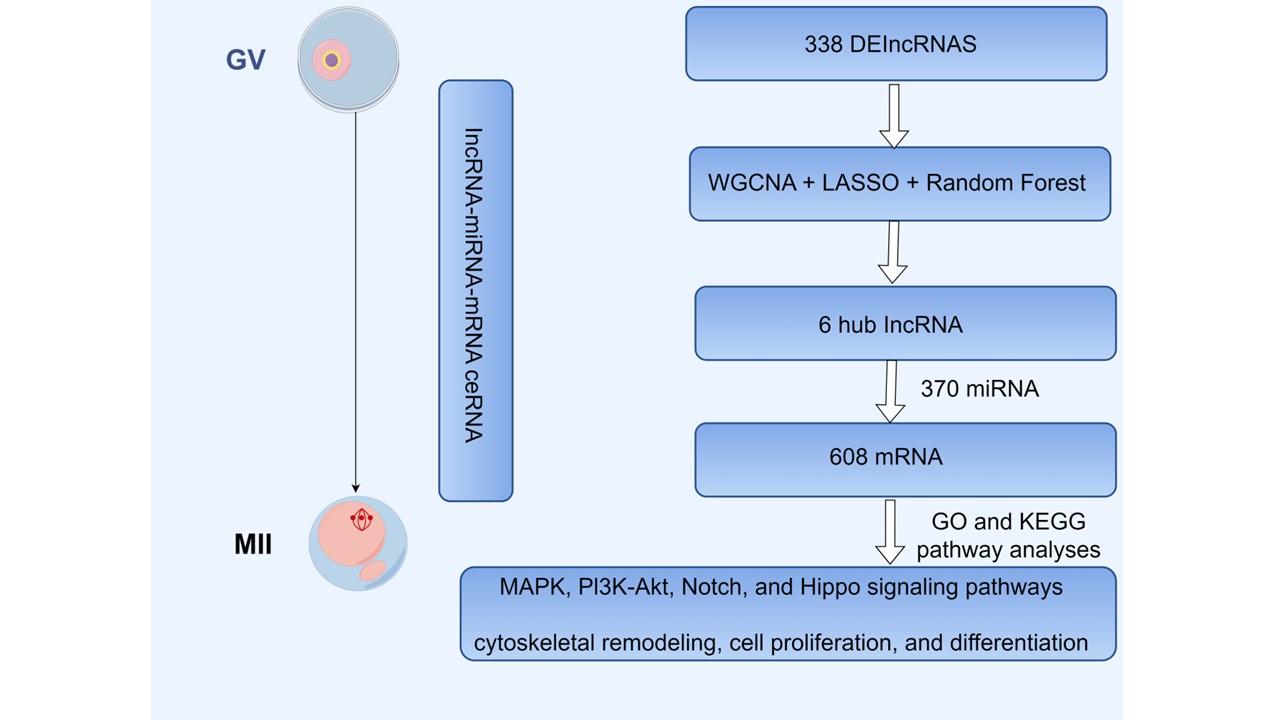
With the increase in maternal age and the impact of environmental stress, the decline in ovarian reserve and oocyte quality has emerged as a primary cause of infertility. Dysfunction of cytoskeletal proteins plays a central role in this process. This study aims to examine the differential expression and regulatory functions of cytoskeleton-associated long non-coding RNAs (lncRNAs) during the development of mouse oocytes at the germinal vesicle (GV) and metaphase II (MII) stages. This study employed bioinformatics analyses and machine learning techniques to analyze publicly accessible data from the Gene Expression Omnibus (GEO) database, which comprised 13 samples of Germinal Vesicle (GV) stage oocytes and 15 samples of Metaphase II (MII) stage oocytes. Differential expression analysis, weighted gene co-expression network analysis (WGCNA), and interaction network construction were performed to screen for lncRNAs closely related to oocyte development. A total of 338 differentially expressed lncRNAs (DE-lncRNAs) with statistical significance were identified, including 136 upregulated and 202 downregulated lncRNAs, indicating their potential roles in the transition from the GV to the MII stage during oocyte development. WGCNA further identified modules strongly correlated with cytoskeletal proteins by integrating these results with the differentially expressed lncRNAs. A total of 47 candidate lncRNAs were shortlisted. Subsequently, LASSO regression and random forest algorithms were applied to identify six key lncRNAs from the candidate set. Combined with miRNA prediction and target gene analysis, a lncRNA-miRNA-mRNA regulatory network was constructed, revealing that these key lncRNAs may indirectly regulate downstream target gene expression through specific miRNAs. Furthermore, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses indicated that these key lncRNAs are primarily involved in cytoskeletal remodeling, cell proliferation, and differentiation, and may play critical roles in follicle structure formation and oocyte development. This study systematically mapped the regulatory network of lncRNAs during oocyte development and elucidated the lncRNA-miRNA-mRNA interactions. The results emphasize the key roles of lncRNAs in cytoskeletal remodeling and oocyte maturation, providing valuable insights for the diagnosis and treatment of ovarian disorders.
References
Barragán, M., Pons, J., Ferrer-Vaquer, A., Cornet-Bartolomé, D., Schweitzer, A., Hubbard, J., Auer, H., Rodolosse, A., & Vassena, R. (2017). The transcriptome of human oocytes is related to age and ovarian reserve. Molecular Human Reproduction, 23(8), 535–548.
Bindea, G., Mlecnik, B., Hackl, H., Charoentong, P., Tosolini, M., Kirilovsky, A., Fridman, W.-H., Pagès, F., Trajanoski, Z., & Galon, J. (2009). ClueGO: A cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics (Oxford, England), 25(8), 1091–1093.
Campbell, H. K., Maiers, J. L., & DeMali, K. A. (2017). Interplay between tight junctions & adherens junctions. Experimental Cell Research, 358(1), 39–44.
Colin, A., Singaravelu, P., Théry, M., Blanchoin, L., & Gueroui, Z. (2018). Actin-network architecture regulates microtubule dynamics. Current Biology : CB, 28(16), 2647-2656.e4.
Ernst, E. H., Nielsen, J., Ipsen, M. B., Villesen, P., & Lykke-Hartmann, K. (2018). Transcriptome analysis of long non-coding RNAs and genes encoding paraspeckle proteins during human ovarian follicle development. Frontiers in Cell and Developmental Biology, 6, 78.
Gardel, M. L., Schneider, I. C., Aratyn-Schaus, Y., & Waterman, C. M. (2010). Mechanical integration of actin and adhesion dynamics in cell migration. Annual Review of Cell and Developmental Biology, 26, 315–333.
Greig, J., & Bulgakova, N. A. (2020). Interplay between actomyosin and E-cadherin dynamics regulates cell shape in the drosophila embryonic epidermis. Journal of Cell Science, 133(15), jcs242321.
Guzmán, A., Hernández-Coronado, C. G., Gutiérrez, C. G., & Rosales-Torres, A. M. (2023). The vascular endothelial growth factor (VEGF) system as a key regulator of ovarian follicle angiogenesis and growth. Molecular Reproduction and Development, 90(4), 201–217.
Hart, R. J. (2016). Physiological Aspects of Female Fertility: Role of the Environment, Modern Lifestyle, and Genetics. Physiological Reviews, 96(3), 873–909.
He, Y.-T., Yang, L.-L., Luo, S.-M., Shen, W., Yin, S., & Sun, Q.-Y. (2019). PAK4 regulates actin and microtubule dynamics during meiotic maturation in mouse oocyte. International Journal of Biological Sciences, 15(11), 2408–2418.
Hu, L.-L., Su, T., Luo, R.-C., Zheng, Y.-H., Huang, J., Zhong, Z.-S., Nie, J., & Zheng, L.-P. (2019). Hippo pathway functions as a downstream effector of AKT signaling to regulate the activation of primordial follicles in mice. Journal of Cellular Physiology, 234(2), 1578–1587.
Hubbard, N., Prasasya, R. D., & Mayo, K. E. (2019). Activation of notch signaling by oocytes and Jag1 in mouse ovarian granulosa cells. Endocrinology, 160(12), 2863–2876.
Irles, P., Elshaer, N., & Piulachs, M.-D. (2016). The notch pathway regulates both the proliferation and differentiation of follicular cells in the panoistic ovary of blattella germanica. Open Biology, 6(1), 150197.
Jiao, J., Shi, B., Wang, T., Fang, Y., Cao, T., Zhou, Y., Wang, X., & Li, D. (2018). Characterization of long non-coding RNA and messenger RNA profiles in follicular fluid from mature and immature ovarian follicles of healthy women and women with polycystic ovary syndrome. Human Reproduction (Oxford, England), 33(9), 1735–1748.
Langfelder, P., & Horvath, S. (2008). WGCNA: An R package for weighted correlation network analysis. BMC Bioinformatics, 9, 559.
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E., & Storey, J. D. (2012). The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics (Oxford, England), 28(6), 882–883.
Li, D., Wang, X., Dang, Y., Zhang, X., Zhao, S., Lu, G., Chan, W.-Y., Leung, P. C. K., & Qin, Y. (2021). lncRNA GCAT1 is involved in premature ovarian insufficiency by regulating p27 translation in GCs via competitive binding to PTBP1. Molecular Therapy - Nucleic Acids, 23, 132–141.
Li, Y., & Tan, Y. (2021). Bioinformatics analysis of ceRNA network related to polycystic ovarian syndrome. Computational and Mathematical Methods in Medicine, 2021, 9988347.
Long, H., Yu, W., Yu, S., Yin, M., Wu, L., Chen, Q., Cai, R., Suo, L., Wang, L., Lyu, Q., & Kuang, Y. (2021). Progesterone affects clinic oocyte yields by coordinating with follicle stimulating hormone via PI3K/AKT and MAPK pathways. Journal of Advanced Research, 33, 189–199.
Mangiavacchi, A., Morelli, G., & Orlando, V. (2023). Behind the scenes: How RNA orchestrates the epigenetic regulation of gene expression. Frontiers in Cell and Developmental Biology, 11, 1123975.
Mao, L., Lou, H., Lou, Y., Wang, N., & Jin, F. (2014). Behaviour of cytoplasmic organelles and cytoskeleton during oocyte maturation. Reproductive Biomedicine Online, 28(3), 284–299.
Mattick, J.S., Amaral, P.P., Carninci, P. et al. (2023). Long non-coding RNAs: Definitions, functions, challenges and recommendations. Nature Reviews. Molecular Cell Biology, 24(6), 430–447.
Mogollón García, H. D., de Andrade Ferrazza, R., Ochoa, J. C., de Athayde, F. F., Vidigal, P. M. P., Wiltbank, M., Kastelic, J. P., Sartori, R., & Ferreira, J. C. P. (2024). Landscape transcriptomic analysis of bovine follicular cells during key phases of ovarian follicular development. Biological Research, 57(1), 76.
Ortega Serrano, P. V., Guzmán, A., Hernández-Coronado, C. G., Castillo-Juárez, H., & Rosales-Torres, A. M. (2016). Reduction in the mRNA expression of sVEGFR1 and sVEGFR2 is associated with the selection of dominant follicle in cows. Reproduction in Domestic Animals = Zuchthygiene, 51(6), 985–991.
Ritchie, M. E., Phipson, B., Wu, D., Hu, Y., Law, C. W., Shi, W., & Smyth, G. K. (2015). Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Research, 43(7), e47.
Rodriguez-Wallberg, K. A., Hao, X., Marklund, A., Johansen, G., Borgström, B., & Lundberg, F. E. (2021). Hot Topics on Fertility Preservation for Women and Girls-Current Research, Knowledge Gaps, and Future Possibilities. Journal of Clinical Medicine, 10(8).
Roeles, J., & Tsiavaliaris, G. (2019). Actin-microtubule interplay coordinates spindle assembly in human oocytes. Nature Communications, 10(1), 4651.
Salmena, L., Poliseno, L., Tay, Y., Kats, L., & Pandolfi, P. P. (2011). A ceRNA hypothesis: The rosetta stone of a hidden RNA language? Cell, 146(3), 353–358.
Shen, M., Li, T., Feng, Y., Wu, P., Serrano, B. R., Barcenas, A. R., Qu, L., & Zhao, W. (2023). Effects of quercetin on granulosa cells from prehierarchical follicles by modulating MAPK signaling pathway in chicken. Poultry Science, 102(7), 102736.
Song, G.-Y., Guo, X.-N., Yao, J., Lu, Z.-N., Xie, J.-H., Wu, F., He, J., Fu, Z.-L., & Han, J. (2023). Differential expression profiles and functional analysis of long non-coding RNAs in calcific aortic valve disease. BMC Cardiovascular Disorders, 23(1), 326.
Wei, L., Xia, H., Liang, Z., Yu, H., Liang, Z., Yang, X., & Li, Y. (2022). Disrupted expression of long non-coding RNAs in the human oocyte: The possible epigenetic culprits leading to recurrent oocyte maturation arrest. Journal of Assisted Reproduction and Genetics, 39(10), 2215–2225.
Wu, T., Dong, J., Fu, J., Kuang, Y., Chen, B., Gu, H., Luo, Y., Gu, R., Zhang, M., Li, W., Dong, X., Sun, X., Sang, Q., & Wang, L. (2022). The mechanism of acentrosomal spindle assembly in human oocytes. Science (New York, N.Y.), 378(6621), eabq7361.
Yamada, T., Kuramitsu, K., Rikitsu, E., Kurita, S., Ikeda, W., & Takai, Y. (2013). Nectin and junctional adhesion molecule are critical cell adhesion molecules for the apico-basal alignment of adherens and tight junctions in epithelial cells. Genes to Cells : Devoted to Molecular & Cellular Mechanisms, 18(11), 985–998.
Yin, X., Wang, P., Yang, T., Li, G., Teng, X., Huang, W., & Yu, H. (2020). Identification of key modules and genes associated with breast cancer prognosis using WGCNA and ceRNA network analysis. Aging, 13(2), 2519–2538.
Zhang, B., & Horvath, S. (2005). A general framework for weighted gene co-expression network analysis. Statistical Applications in Genetics and Molecular Biology, 4, Article17.
Zhang, L., Zou, J., Wang, Z., & Li, L. (2023). A subpathway and target gene cluster-based approach uncovers lncRNAs associated with human primordial follicle activation. International Journal of Molecular Sciences, 24(13).
Zhang, Y., Wan, X., Wang, H.-H., Pan, M.-H., Pan, Z.-N., & Sun, S.-C. (2019). RAB35 depletion affects spindle formation and actin-based spindle migration in mouse oocyte meiosis. Molecular Human Reproduction, 25(7), 359–372.
Downloads
Published
Issue
Section
License
Copyright (c) 2025 Malaysian Journal of Science

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
Transfer of Copyrights
- In the event of publication of the manuscript entitled [INSERT MANUSCRIPT TITLE AND REF NO.] in the Malaysian Journal of Science, I hereby transfer copyrights of the manuscript title, abstract and contents to the Malaysian Journal of Science and the Faculty of Science, University of Malaya (as the publisher) for the full legal term of copyright and any renewals thereof throughout the world in any format, and any media for communication.
Conditions of Publication
- I hereby state that this manuscript to be published is an original work, unpublished in any form prior and I have obtained the necessary permission for the reproduction (or am the owner) of any images, illustrations, tables, charts, figures, maps, photographs and other visual materials of whom the copyrights is owned by a third party.
- This manuscript contains no statements that are contradictory to the relevant local and international laws or that infringes on the rights of others.
- I agree to indemnify the Malaysian Journal of Science and the Faculty of Science, University of Malaya (as the publisher) in the event of any claims that arise in regards to the above conditions and assume full liability on the published manuscript.
Reviewer’s Responsibilities
- Reviewers must treat the manuscripts received for reviewing process as confidential. It must not be shown or discussed with others without the authorization from the editor of MJS.
- Reviewers assigned must not have conflicts of interest with respect to the original work, the authors of the article or the research funding.
- Reviewers should judge or evaluate the manuscripts objective as possible. The feedback from the reviewers should be express clearly with supporting arguments.
- If the assigned reviewer considers themselves not able to complete the review of the manuscript, they must communicate with the editor, so that the manuscript could be sent to another suitable reviewer.
Copyright: Rights of the Author(s)
- Effective 2007, it will become the policy of the Malaysian Journal of Science (published by the Faculty of Science, University of Malaya) to obtain copyrights of all manuscripts published. This is to facilitate:
- Protection against copyright infringement of the manuscript through copyright breaches or piracy.
- Timely handling of reproduction requests from authorized third parties that are addressed directly to the Faculty of Science, University of Malaya.
- As the author, you may publish the fore-mentioned manuscript, whole or any part thereof, provided acknowledgement regarding copyright notice and reference to first publication in the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers) are given. You may produce copies of your manuscript, whole or any part thereof, for teaching purposes or to be provided, on individual basis, to fellow researchers.
- You may include the fore-mentioned manuscript, whole or any part thereof, electronically on a secure network at your affiliated institution, provided acknowledgement regarding copyright notice and reference to first publication in the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers) are given.
- You may include the fore-mentioned manuscript, whole or any part thereof, on the World Wide Web, provided acknowledgement regarding copyright notice and reference to first publication in the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers) are given.
- In the event that your manuscript, whole or any part thereof, has been requested to be reproduced, for any purpose or in any form approved by the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers), you will be informed. It is requested that any changes to your contact details (especially e-mail addresses) are made known.
Copyright: Role and responsibility of the Author(s)
- In the event of the manuscript to be published in the Malaysian Journal of Science contains materials copyrighted to others prior, it is the responsibility of current author(s) to obtain written permission from the copyright owner or owners.
- This written permission should be submitted with the proof-copy of the manuscript to be published in the Malaysian Journal of Science
Licensing Policy
Malaysian Journal of Science is an open-access journal that follows the Creative Commons Attribution-Non-commercial 4.0 International License (CC BY-NC 4.0)
CC BY – NC 4.0: Under this licence, the reusers to distribute, remix, alter, and build upon the content in any media or format for non-commercial purposes only, as long as proper acknowledgement is given to the authors of the original work. Please take the time to read the whole licence agreement (https://creativecommons.org/licenses/by-nc/4.0/legalcode ).





