Bioinformatics Analysis of Potential Biomarkers for Lupus Nephritis
DOI:
https://doi.org/10.22452/mjs.vol44sp1.8Keywords:
Lupus Nephritis, bioinformatics, proteomics, biomarkerAbstract
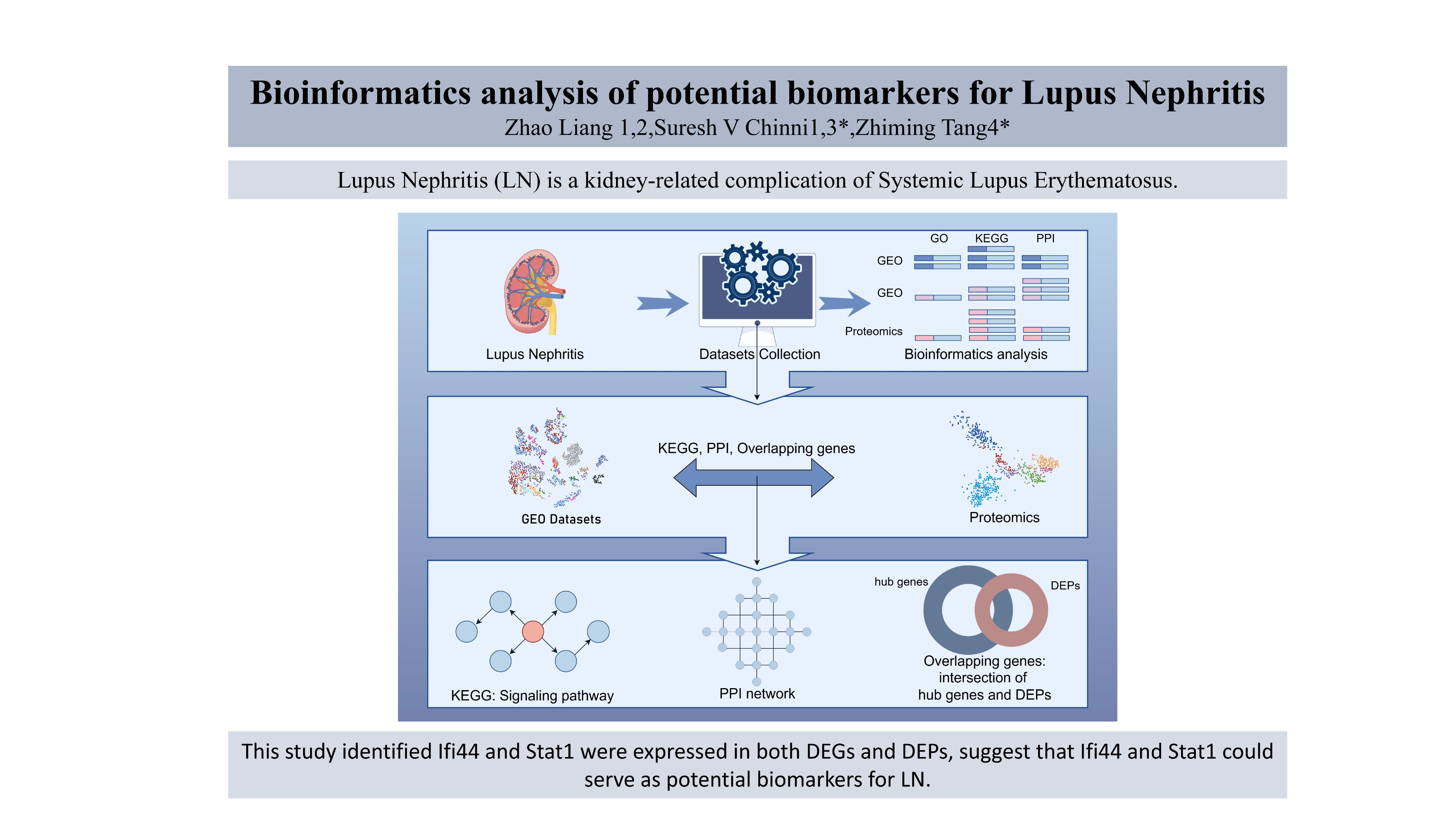
Lupus Nephritis (LN) is a complication of Systemic Lupus Erythematosus affecting the kidney. The purpose of this study was to identify signalling pathways and hub genes involved in the pathogenesis of LN. Methods: The mRNA expression profiles of LN were obtained from the Gene Expression Omnibus database, and differentially expressed genes (DEGs) were identified using the online tool GEO2R. Enrichment analysis was conducted in DAVID. The Protein-Protein Interaction network of DEGs was constructed in STRING, and hub genes were identified with Cytoscape. The hub genes were validated using differentially expressed proteins (DEPs) from proteomics data to identify potential biomarkers for LN. Results: A total of 138 DEGs were identified, primarily associated with immune response, neutrophil chemotaxis, and antimicrobial humoral immunity. In KEGG analysis, the NOD-like receptor signalling pathway and the Cytokine-cytokine receptor interaction pathway were mainly involved. Nine hub genes of LN, including Ifit1, Ifit3, Ifih1, Ifi44, Irf7, Irf9, Oasl1, Stat1, and Usp18 were identified. Conclusion: Ifi44 and Stat1 were expressed in both DEGs and DEPs. Ifi44 and Stat1 may be potential biomarkers and therapeutic targets for LN.
References
Arazi, A., Rao, D. A., Berthier, C. C., Davidson, A., Liu, Y., et al. (2019). The immune cell landscape in kidneys of patients with lupus nephritis. Nature Immunology, 20(7), 902–914.
Arora, P., Malik, M., Sachdeva, R., Saxena, L., Das, J., Ramachandran, V. G., et al. (2017). Innate and humoral recognition of the products of cell death: Differential antigenicity and immunogenicity in lupus. Clinical and Experimental Immunology, 187(3), 353–368.
Berthier, C. C., Bethunaickan, R., Gonzalez-Rivera, T., Nair, V., Ramanujam, M., Zhang, W., et al. (2012). Cross-species transcriptional network analysis defines shared inflammatory responses in murine and human lupus nephritis. Journal of Immunology, 189(2), 988–1001.
Bethunaickan, R., Berthier, C. C., Ramanujam, M., Sahu, R., Zhang, W., Sun, Y., et al. (2011). A unique hybrid renal mononuclear phagocyte activation phenotype in murine systemic lupus erythematosus nephritis. Journal of Immunology, 186(8), 4994–5003.
Cai, Z., Zhang, S., Wu, P., Ren, Q., Wei, P., Hong, M., et al. (2021). A novel potential target of IL-35–regulated JAK/STAT signaling pathway in lupus nephritis. Clinical and Translational Medicine, 11(2), e309.
Chen, J., Li, M., Shang, S., Cheng, L., Tang, Z., & Huang, C. (2024). LncRNA XIST/miR-381-3P/STAT1 axis as a potential biomarker for lupus nephritis. Lupus, 33(11), 1176–1191.
Coit, P., Renauer, P., Jeffries, M. A., Merrill, J. T., McCune, W. J., Maksimowicz-McKinnon, K., et al. (2015). Renal involvement in lupus is characterized by unique DNA methylation changes in naïve CD4+ T cells. Journal of Autoimmunity, 61, 29–35.
Conti, F., Spinelli, F. R., Alessandri, C., & Valesini, G. (2011). Toll-like receptors and lupus nephritis. Clinical Reviews in Allergy & Immunology, 41(3), 192–198.
Davidson, A., Bethunaickan, R., Berthier, C., Sahu, R., Zhang, W., & Kretzler, M. (2015). Molecular studies of lupus nephritis kidneys. Immunological Research, 63(1–3), 187–196.
Deng, Y., Zheng, Y., Li, D., Hong, Q., Zhang, M., Li, Q., et al. (2021). Expression characteristics of interferon-stimulated genes and possible regulatory mechanisms in lupus patients using transcriptomics analyses. EBioMedicine, 70, 103477.
Elkhateeb, E., Tag-El-Din-Hassan, H. T., Sasaki, N., Torigoe, D., Morimatsu, M., & Agui, T. (2016). The role of mouse 2′,5′-oligoadenylate synthetase 1 paralogs. Infection, Genetics and Evolution, 45, 393–401.
Funabiki, M., Kato, H., Miyachi, Y., Toki, H., Motegi, H., Inoue, M., et al. (2014). Autoimmune disorders associated with gain of function of the intracellular sensor MDA5. Immunity, 40(2), 199–212.
Gardet, A., Chou, W. C., Reynolds, T. L., Velez, D. B., Fu, K., Czerkowicz, J. M., et al. (2016). Pristane-accelerated autoimmune disease in (SWR × NZB) F1 mice leads to prominent tubulointerstitial inflammation and human lupus nephritis-like fibrosis. PLOS ONE, 11(10), e0164423.
Gibbert, K., François, S., Sigmund, A. M., Harper, M. S., Barrett, B. S., Kirschning, C. J., et al. (2014). Friend retrovirus drives cytotoxic effectors through Toll-like receptor 3. Retrovirology, 11, 126.
Giannico, G., & Fogo, A. B. (2013). Lupus nephritis: Is the kidney biopsy currently necessary in the management of lupus nephritis? Clinical Journal of the American Society of Nephrology, 8(1), 138–145.
Huang, D. W., Sherman, B. T., & Lempicki, R. A. (2009). Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols, 4(1), 44–57.
Hu, W., Niu, G., Li, H., Gao, H., Kang, R., Chen, X., et al. (2016). The association between expression of IFIT1 in podocytes of MRL/lpr mice and the renal pathological changes it causes. Oncotarget, 7(47), 76464–76470.
Ikeda, K., Hayakawa, K., Fujishiro, M., Kawasaki, M., Hirai, T., Tsushima, H., et al. (2017). JAK inhibitor has the amelioration effect in lupus-prone mice: The involvement of IFN signature gene downregulation. BMC Immunology, 18, 41.
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M., & Tanabe, M. (2021). KEGG: Integrating viruses and cellular organisms. Nucleic Acids Research, 49(D1), D545–D551.
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M., & Ishiguro-Watanabe, M. (2023). KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Research, 51(D1), D587–D592.
Lai, J. H., Wu, D. W., Huang, C. Y., Hung, L. F., Wu, C. H., & Ho, L. J. (2024). USP18 induction regulates immunometabolism to attenuate M1 signal-polarized macrophages and enhance IL-4–polarized macrophages in systemic lupus erythematosus. Clinical Immunology, 265, 110285.
Lech, M., & Anders, H. J. (2013). The pathogenesis of lupus nephritis. Journal of the American Society of Nephrology, 24(9), 1357–1366.
Lee, M. S., Park, C. H., Jeong, Y. H., Kim, Y. J., & Ha, S. J. (2013). Negative regulation of type I IFN expression by OASL1 permits chronic viral infection and CD8+ T-cell exhaustion. PLoS Pathogens, 9(7), e1003478.
Liao, Z., Ye, Z., Xue, Z., Wu, L., Ouyang, Y., Yao, C., et al. (2019). Identification of renal long non-coding RNA RP11-2B6.2 as a positive regulator of type I interferon signaling pathway in lupus nephritis. Frontiers in Immunology, 10, 975.
Lin, W., Lin, Z., Lin, X., Peng, Z., Liang, X., & Wei, S. (2023). Integrated analysis of hub genes, immune infiltration, and potential therapeutic agents related to lupus nephritis. Lupus, 32(5), 633–643.
Lu, M. C., Lai, N. S., Chen, H. C., Yu, H. C., Huang, K. Y., Tung, C. H., et al. (2012). Decreased microRNA-145 and increased miR-224 expression in T cells from patients with systemic lupus erythematosus. Clinical and Experimental Immunology, 171(1), 91–99.
Mistry, P., Nakabo, S., O’Neil, L., Goel, R. R., Jiang, K., Carmona-Rivera, C., et al. (2019). Transcriptomic, epigenetic, and functional analyses implicate neutrophil diversity in the pathogenesis of systemic lupus erythematosus. Proceedings of the National Academy of Sciences, 116(50), 25222–25228.
Miyagawa, F., Tagaya, Y., Ozato, K., & Asada, H. (2016). Essential requirement for IFN regulatory factor 7 in autoantibody production but not development of nephritis in murine lupus. Journal of Immunology, 197(6), 2167–2176.
Mok, A., Solomon, O., Nayak, R. R., Coit, P., Quach, H. L., Nititham, J., et al. (2016). Genome-wide profiling identifies associations between lupus nephritis and differential methylation of genes regulating tissue hypoxia and type 1 interferon responses. Lupus Science & Medicine, 3(1), e000183.
Mou, L., Zhang, F., Liu, X., Lu, Y., Yue, M., Lai, Y., et al. (2024). Integrative analysis of COL6A3 in lupus nephritis: Insights from single-cell transcriptomics and proteomics. Frontiers in Immunology, 15, 1309447.
Nicolaou, O., Sokratous, K., Makowska, Z., Morell, M., De Groof, A., Montigny, P., et al. (2020). Proteomic analysis in lupus mice identifies Coronin-1A as a potential biomarker for lupus nephritis. Arthritis Research & Therapy, 22, 147.
Pang, J., Huang, P., Huang, H., Ma, J., He, L., Lin, X., et al. (2025). Molecular mechanism and role of miRNA-155 in podocyte apoptosis in lupus nephritis. International Journal of Biological Macromolecules, 304, 140810.
Parikh, S. V., Almaani, S., Brodsky, S., & Rovin, B. H. (2020). Update on lupus nephritis: Core curriculum 2020. American Journal of Kidney Diseases, 76(2), 265–281.
Paul, P., Antonydhason, V., Gopal, J., Haga, S. W., Hasan, N., & Oh, J. W. (2020).
Bioinformatics for renal and urinary proteomics: Call for aggrandization. International Journal of Molecular Sciences, 21(3), 961.
Qing, X., Chinenov, Y., Redecha, P., Madaio, M., Roelofs, J. J. T. H., Farber, G., et al. (2018). iRhom2 promotes lupus nephritis through TNF-α and EGFR signaling. Journal of Clinical Investigation, 128(4), 1397–1412.
Saxena, R., Mahajan, T., & Mohan, C. (2011). Lupus nephritis: Current update. Arthritis Research & Therapy, 13(5), 240.
Shen, L., Lan, L., Zhu, T., Chen, H., Gu, H., Wang, C., et al. (2021). Identification and validation of IFI44 as a key biomarker in lupus nephritis. Frontiers in Medicine, 8, 762848.
Shen, M., Duan, C., Xie, C., Wang, H., Li, Z., Li, B., et al. (2022). Identification of key interferon-stimulated genes for indicating the condition of patients with systemic lupus erythematosus. Frontiers in Immunology, 13, 962393.
Szklarczyk, D., Gable, A. L., Nastou, K. C., Lyon, D., Kirsch, R., Pyysalo, S., et al. (2021). The STRING database in 2021: Customizable protein–protein networks and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Research, 49(D1), D605–D612.
Tang, D., Chen, M., Huang, X., Zhang, G., Zeng, L., Zhang, G., et al. (2023). SRplot: A free online platform for data visualization and graphing. PLOS ONE, 18(11), e0294236.
Tang, Y., Zhang, Y., Li, X., Xu, R., Ji, Y., Liu, J., et al. (2023). Immune landscape and the key role of APOE+ monocytes in lupus nephritis under single-cell and spatial transcriptional analysis. Clinical and Translational Medicine, 13(4), e1237.
The Gene Ontology Consortium, Carbon, S., Douglass, E., Good, B. M., Unni, D. R., Harris, N. L., et al. (2021). The Gene Ontology resource: Enriching a GOld mine. Nucleic Acids Research, 49(D1), D325–D334.
Thibault, D. L., Chu, A. D., Graham, K. L., Balboni, I., Lee, L. Y., Kohlmoos, C., et al. (2008). IRF9 and STAT1 are required for IgG autoantibody production and B cell expression of TLR7 in mice. Journal of Clinical Investigation, 118(4), 1417–1426.
Wen, Q., Wang, C., Chen, D., Luo, N., Fan, J., Zhou, Y., et al. (2024). Proteomics-based identification of therapeutic targets of artesunate in a lupus nephritis MRL/lpr mouse model. Journal of Proteome Research, 23(4), 1150–1162.
Yiu, G., Rasmussen, T. K., Ajami, B., Haddon, D. J., Chu, A. D., Tangsombatvisit, S., et al. (2016). Development of Th17-associated interstitial kidney inflammation in lupus-prone mice lacking STAT1. Arthritis & Rheumatology, 68(5), 1233–1244.
Yu, C., Li, P., Dang, X., Zhang, X., Mao, Y., & Chen, X. (2022). Lupus nephritis: New progress in diagnosis and treatment. Journal of Autoimmunity, 132, 102871.
Zhang, H., Liu, L., & Li, L. (2018). Lentivirus-mediated knockdown of FcγRI attenuates lupus nephritis via inhibition of NF-κB and NLRP3 inflammasome activation in MRL/lpr mice. Journal of Pharmacological Sciences, 137(4), 342–349.
Zhao, J., Wang, H., Dai, C., Wang, H., Zhang, H., Huang, Y., et al. (2013). P2X7 blockade attenuates murine lupus nephritis by inhibiting activation of the NLRP3/ASC/caspase-1 pathway. Arthritis & Rheumatism, 65(12), 3176–3185.
Zheng, C., Shang, F., Cheng, R., & Bai, Y. (2024). STAT1 aggravates kidney injury by NLRP3 signaling in MRL/lpr mice. Journal of Molecular Histology, 55(4), 555–566.
Zhou, G., Soufan, O., Ewald, J., Hancock, R. E. W., Basu, N., & Xia, J. (2019).
NetworkAnalyst 3.0: A visual analytics platform for comprehensive gene expression profiling and meta-analysis. Nucleic Acids Research, 47(W1), W234–W241.
Downloads
Published
Issue
Section
License
Copyright (c) 2025 Malaysian Journal of Science

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
Transfer of Copyrights
- In the event of publication of the manuscript entitled [INSERT MANUSCRIPT TITLE AND REF NO.] in the Malaysian Journal of Science, I hereby transfer copyrights of the manuscript title, abstract and contents to the Malaysian Journal of Science and the Faculty of Science, University of Malaya (as the publisher) for the full legal term of copyright and any renewals thereof throughout the world in any format, and any media for communication.
Conditions of Publication
- I hereby state that this manuscript to be published is an original work, unpublished in any form prior and I have obtained the necessary permission for the reproduction (or am the owner) of any images, illustrations, tables, charts, figures, maps, photographs and other visual materials of whom the copyrights is owned by a third party.
- This manuscript contains no statements that are contradictory to the relevant local and international laws or that infringes on the rights of others.
- I agree to indemnify the Malaysian Journal of Science and the Faculty of Science, University of Malaya (as the publisher) in the event of any claims that arise in regards to the above conditions and assume full liability on the published manuscript.
Reviewer’s Responsibilities
- Reviewers must treat the manuscripts received for reviewing process as confidential. It must not be shown or discussed with others without the authorization from the editor of MJS.
- Reviewers assigned must not have conflicts of interest with respect to the original work, the authors of the article or the research funding.
- Reviewers should judge or evaluate the manuscripts objective as possible. The feedback from the reviewers should be express clearly with supporting arguments.
- If the assigned reviewer considers themselves not able to complete the review of the manuscript, they must communicate with the editor, so that the manuscript could be sent to another suitable reviewer.
Copyright: Rights of the Author(s)
- Effective 2007, it will become the policy of the Malaysian Journal of Science (published by the Faculty of Science, University of Malaya) to obtain copyrights of all manuscripts published. This is to facilitate:
- Protection against copyright infringement of the manuscript through copyright breaches or piracy.
- Timely handling of reproduction requests from authorized third parties that are addressed directly to the Faculty of Science, University of Malaya.
- As the author, you may publish the fore-mentioned manuscript, whole or any part thereof, provided acknowledgement regarding copyright notice and reference to first publication in the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers) are given. You may produce copies of your manuscript, whole or any part thereof, for teaching purposes or to be provided, on individual basis, to fellow researchers.
- You may include the fore-mentioned manuscript, whole or any part thereof, electronically on a secure network at your affiliated institution, provided acknowledgement regarding copyright notice and reference to first publication in the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers) are given.
- You may include the fore-mentioned manuscript, whole or any part thereof, on the World Wide Web, provided acknowledgement regarding copyright notice and reference to first publication in the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers) are given.
- In the event that your manuscript, whole or any part thereof, has been requested to be reproduced, for any purpose or in any form approved by the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers), you will be informed. It is requested that any changes to your contact details (especially e-mail addresses) are made known.
Copyright: Role and responsibility of the Author(s)
- In the event of the manuscript to be published in the Malaysian Journal of Science contains materials copyrighted to others prior, it is the responsibility of current author(s) to obtain written permission from the copyright owner or owners.
- This written permission should be submitted with the proof-copy of the manuscript to be published in the Malaysian Journal of Science
Licensing Policy
Malaysian Journal of Science is an open-access journal that follows the Creative Commons Attribution-Non-commercial 4.0 International License (CC BY-NC 4.0)
CC BY – NC 4.0: Under this licence, the reusers to distribute, remix, alter, and build upon the content in any media or format for non-commercial purposes only, as long as proper acknowledgement is given to the authors of the original work. Please take the time to read the whole licence agreement (https://creativecommons.org/licenses/by-nc/4.0/legalcode ).





